Lock-and-key model
strong>Lock-and-key model n., [lɑk ænd ki ˈmɑdl̩] Definition: a model for enzyme-substrate interaction
Table of Contents

Lock-and-key model Definition
Lock-and-key model is a model for enzyme-substrate interaction suggesting that the enzyme and the substrate possess specific complementary geometric shapes that fit exactly into one another. In this model, enzymes are depicted as highly specific. They must bind to specific substrates before they catalyze chemical reactions . The term is a pivotal concept in enzymology to elucidate the intricate interaction between enzymes and substrates at the molecular level. In the lock-and-key model, the enzyme-substrate interaction suggests that the enzyme and the substrate possess specific complementary geometric shapes that fit exactly into one another. Like a key into a lock , only the correct size and shape of the substrate ( the key ) would fit into the active site ( the keyhole ) of the enzyme ( the lock ).
Compare: Induced fit model See also: enzyme , active site , substrate
Lock-and-key vs. Induced Fit Model
At present, two models attempt to explain enzyme-substrate specificity; one of which is the lock-and-key model , and the other is the Induced fit model . The lock and key model theory was first postulated by Emil Fischer in 1894. The lock-and-key enzyme action proposes the high specificity of enzymes. However, it does not explain the stabilization of the transition state that the enzymes achieve. The induced fit model (proposed by Daniel Koshland in 1958) suggests that the active site continues to change until the substrate is completely bound to the active site of the enzyme, at which point the final shape and charge are determined. Unlike the lock-and-key model, the induced fit model shows that enzymes are rather flexible structures. Nevertheless, Fischer’s Lock and Key theory laid an important foundation for subsequent research, such as during the refinement of the enzyme-substrate complex mechanism, as ascribed in the induced fit model. The lock-and-key hypothesis has opened ideas where enzyme action is not merely catalytic but incorporates a rather complex process in how they interact with the correct substrates with precision.
Key Components
Components of the lock and key model:
- Enzyme : the enzyme structure is a three-dimensional protein configuration, with an active site from where the substrate binds.
- Substrate : often an organic molecule, a substrate possesses a structural feature that complements the geometry of the enzyme’s active site.
In the lock and key model, both the enzymes and the substrates facilitate the formation of a complex that lowers the activation energy needed for a chemical transformation to occur. Such reduction in the activation energy allows the chemical reaction to proceed at a relatively faster rate, making enzymes crucial in various biological and molecular processes.
Lock-and-key Model Examples
Some of the common examples that are often discussed in the context of the Lock and Key Model are as follows:
- Enzyme lactate dehydrogenase with a specific active site for its substrates, pyruvate and lactate. The complex facilitates the interconversion of pyruvate and lactate during anaerobic respiration
- Enzyme carbonic anhydrase with a specific active site for the substrates carbon dioxide and water. The complex facilitates the hydration of carbon dioxide, forming bicarbonate
- Enzyme lysozyme binding with a bacterial cell wall peptidoglycan, which is a vital immune function
Choose the best answer.
Send Your Results (Optional)
- Aryal, S. and Karki, P. (2023). “Lock and Key Model- Mode of Action of Enzymes”. Microbenotes.com. https://microbenotes.com/lock-and-key-model-mode-of-action-of-enzymes/
- Farhana, A., & Lappin, S. L. (2023, May). Biochemistry, Lactate Dehydrogenase . Nih.gov; StatPearls Publishing. https://www.ncbi.nlm.nih.gov/books/NBK557536/
©BiologyOnline.com. Content provided and moderated by Biology Online Editors.
Last updated on January 11th, 2024
You will also like...
New zealand’s unique fauna.
Meet some of New Zealand's unique fauna, including endemic insects, frogs, reptiles, birds, and mammals, and investigate..
Growth and Development of a Human Baby
Upon fertilization, a zygote forms and develops into an embryo. This tutorial elaborates on the growth and development f..
Still Water Animals
Animals living in aquatic habitats have diversified and evolved through time. They eventually occupy ecological niches a..
Temperature Regulation in Animals
This tutorial elucidates body temperature regulation. Know the details here to learn how the body sets the body temperat..
Lights’ Effect on Growth
This tutorial elaborates on the effect of light on plant growth. It describes how different plants require different amo..
Stems primarily provide plants structural support. This tutorial includes lectures on the external form of a woody twig ..
Related Articles...
No related articles found

Microbe Notes
Lock and Key Model- Mode of Action of Enzymes
Enzymes are biological catalysts. These are commonly proteins but also include RNA (ribozymes) molecules that catalyze chemical reactions by lowering the activation energy of a reaction. These are known to speed up the rate of a reaction millions of times faster than the reaction without enzymes. Nearly all biological reactions require enzymes to transform substrate into products. The substrate is the reactant molecule upon which enzymes act during a chemical reaction, and products are the substances formed as a result of a chemical reaction. A single reactant molecule can decompose to give multiple products. Similarly, two reactants can enter into a reaction to yield products. These are reusable even after the completion of the reaction. Chemical properties such as charge and pH are vital in enzymatic reactions.
Binding between enzymes and reactant molecules takes place in such a way that chemical bond-breaking and bond-forming processes occur more readily. Meanwhile, no change in ∆the G value of a reaction takes place, thereby not altering the energy-releasing or energy-absorbing process of the reaction. However, it lowers the energy of the transition state, the topmost unstable state where the activated complex is formed from reactants that later give products.
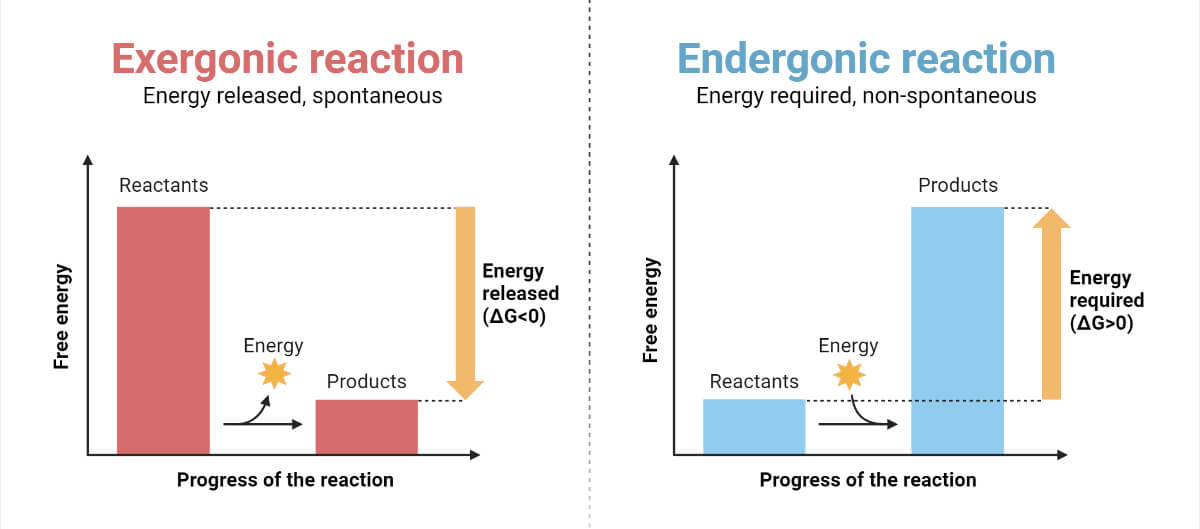
Table of Contents
Interesting Science Videos
Enzyme’s Active site and Substrate Specificity
Enzymes are relatively larger than the substrates, whose only a small fraction is involved in catalysis by reducing chemical activation energy, also known as the catalytic site, and the other portion for binding with the substrate and orienting them also known as the binding site. The catalytic site and binding site altogether form the active site of an enzyme. Usually, there are two active sites in an enzyme.
- The active site of enzymes is a cleft portion, composed of a small number of a unique combination of amino acid residues, usually three to four in number, which make up only ~10-20% of the volume of an enzyme.
- The remaining amino acids are used to maintain tertiary structure by proper scaffold folding through non-covalent interactions.
- Non-covalent interaction between enzyme and substrate in correct orientation favors their reaction. These interactions include hydrogen bonds , hydrophobic bonds, ionic interactions, and Van der Waal’s interactions.
- However, transient covalent bonds between enzymes and substrates are also formed during the time of reaction.
- Side chains of amino acids play an important role in highly specific three-dimensional conformation at the level of the active site. These are large or small, hydrophilic or hydrophobic, acidic or basic.
- The specific shape, size, and chemical behavior of enzymes are determined by the nature of amino acids and their 3D space in the active site.
Specificity is a distinctive feature of enzymes where they have a unique ability to choose an exact substrate from a group of similar chemical molecules. Their specificity towards their substrate varies to a different extent. These are of different types, namely: Bond specificity, Group specificity. Substrate specificity, Stereospecificity, Geometrical specificity, and Co-factor specificity.
Substrate specificity is also k/a absolute specificity for the enzyme’s specificity towards one substrate and one reaction. For e.g., Lactase acts on the B-1-4 glycosidic linkage of lactose to yield galactose and glucose. The restrictive nature of enzymes towards the choice of substrate can be attributed to the enzymatic activity of two oxidoreductase enzymes. Alcohol dehydrogenase uses its substrate alcohol while lactic acid dehydrogenase act on lactic acid. Although these two enzymes function with the mechanism of oxidation and reduction reaction, their substrates can’t be used interchangeably. This is because the different structure of each substrate prevents their fitting into the active site of the alternative enzyme.
In most cases, cofactors, the non-protein molecules, are required to ensure an efficient enzyme-facilitated chemical reaction. These function to bind with enzymes via either ionic interaction or covalent interactions. Metal ions (such as minerals) and co-enzymes (vitamin derivatives) are cofactors.
Lock and Key Model
A German scientist, Emil Fischer postulated the lock and key model in 1894 to explain the enzyme’s mode of action. Fischer’s theory hypothesized that enzymes exhibit a high degree of specificity towards the substrate. This model assumes that the active site of the enzyme and the substrate fit perfectly into one another such that each possesses specific predetermined complementary geometric shapes and sizes. Thus, the shape of the enzyme and substrate do not influence each other. This specificity is analog to the lock and key model, where the lock is the enzyme, and the key is the substrate. In certain circumstances, if a second substrate similar in shape and size to the primary substrate is made to bind to the enzyme, this second substrate also fits in the active site too.
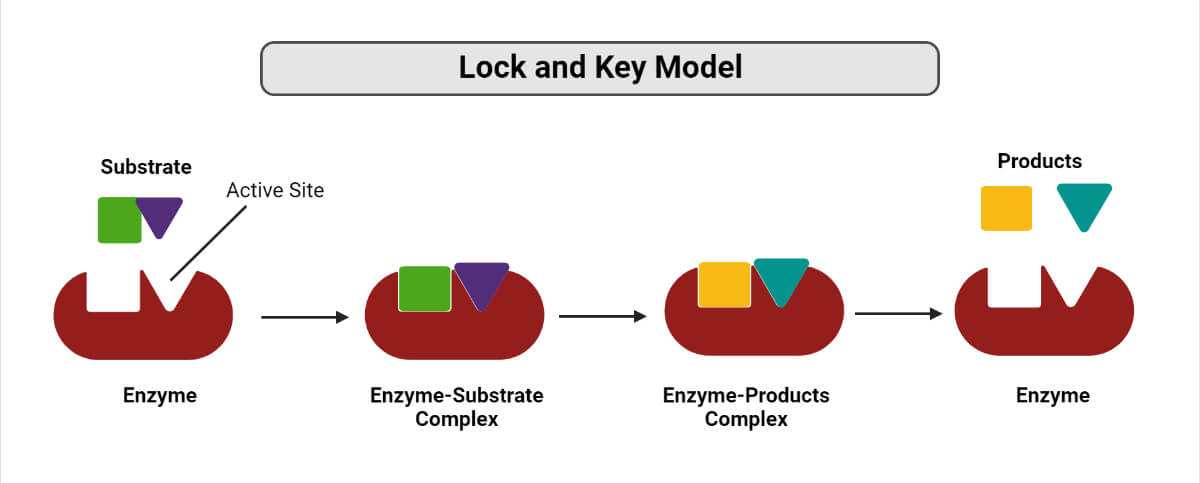
How does Lock and Key Model work?
- Binding of the substrate(s) to the enzyme at their active site takes place, thereby forming an enzyme-substrate complex.
- Enzymes catalyze the chemical reaction to take place, which can either be a synthesis reaction (favors bond formation) or a decomposition reaction (favors bond breakage).
- As a result, the formation of one or more products takes place, and the enzymes are released for their reuse in the next reaction.
Limitations of Lock and Key Model
- It doesn’t explain how the enzyme-substrate complex is stabilized in the transition state.
- This model supposes the enzyme is a rigid structure whose shape does not change upon binding with a suitable substrate. However, this is not supported by recent research, which states that there is a change in conformation of the active site of the enzyme upon binding of substrate.
- It does not describe the condition for binding multiple substrates to the enzyme.
Later, it was found that enzyme specificity toward one substrate is not always true. Although there are enzymes that specifically bind with only one substrate, there are also enzymes that exhibit broad specificity towards different but similarly structured substrates, such as lipase that can bind to different types of lipids. Similarly, proteases such as trypsin and chymotrypsin can degrade multiple types of proteins. Thus, the lock and key model is flawed, and the induced fit model was introduced to give a more refined view of the mode of enzymatic action.
- Blanco, A., & Blanco, G. (2017). Medical Biochemistry. Academic Press. https://www.khanacademy.org/science/ap-biology/cellular-energetics/enzyme-structure-and-catalysis/a/enzymes-and-the-active-site
- https://www.biologyonline.com/dictionary/substrate-specificity
- https://www.britannica.com/science/protein/The-mechanism-of-enzymatic-action
- https://www.biologyonline.com/dictionary/lock-and-key-model
- https://study.com/learn/lesson/lock-key-model-vs-induced-fit-model.html
- https://chem.libretexts.org/Bookshelves/Organic_Chemistry/Organic_Chemistry_(McMurry)/26%3A_Biomolecules-_Amino_Acids_Peptides_and_Proteins/26.11%3A_Enzymes_and_Coenzymes
- https://en.wikibooks.org/wiki/Structural_Biochemistry/Protein_function/Lock_and_Key
- https://ib.bioninja.com.au/higher-level/topic-8-metabolism-cell/untitled-6/models-of-action.html
About Author
Prakriti Karki
Leave a Comment Cancel reply
Save my name, email, and website in this browser for the next time I comment.
Molecular Recognition: Lock-and-Key, Induced Fit, and Conformational Selection
- Reference work entry
- pp 1584–1588
- Cite this reference work entry

- Todd Holyoak 2
Fluctuation fit ; Population selection ; Population shift ; Preexisting equilibrium ; Selected fit
In the most general sense, molecular recognition corresponds to the mechanism by which two or more molecules come together to form a specific complex. These types of specific molecular interactions span biology and include processes as diverse as enzyme catalysis, antibody–antigen recognition, protein synthesis, and transcriptional regulation to name but a few. As a consequence of the universal importance of molecular recognition in biological function, understanding how molecules specifically recognize and interact with one another is of fundamental importance.
Introduction
Early studies on the kinetics and specificity of enzyme-catalyzed reactions led to the conclusion that substrates combine with the enzyme at a specific active site location on the enzyme surface. This understanding generated new questions as to how enzymes could carry out chemical transformations...
This is a preview of subscription content, log in via an institution to check access.
Access this chapter
Subscribe and save.
- Get 10 units per month
- Download Article/Chapter or eBook
- 1 Unit = 1 Article or 1 Chapter
- Cancel anytime
- Available as PDF
- Read on any device
- Instant download
- Own it forever
- Available as EPUB and PDF
- Durable hardcover edition
- Dispatched in 3 to 5 business days
- Free shipping worldwide - see info
Tax calculation will be finalised at checkout
Purchases are for personal use only
Institutional subscriptions
Beach H, Cole R, Gill ML, Loria JP. Conservation of mus-ms enzyme motions in the apo- and substrate-mimicked state. J Am Chem Soc. 2005;127(25):9167–76.
CAS PubMed Google Scholar
Csermely P, Palotai R, Nussinov R. Induced fit, conformational selection and independent dynamic segments: an extended view of binding events. Trends Biochem Sci. 2010;35(10):539–46.
CAS PubMed Central PubMed Google Scholar
Dixon M, Webb EC. Enzymes. New York: Academic; 1979.
Google Scholar
Fischer E. The influence of configuration on enzyme activity. Dtsch Chem Ges. 1894;27:2984–93 (Translated from German).
Gerstein M, Lesk AM, Chothia C. Structural mechanisms for domain movements in proteins. Biochemistry. 1994;33(22):6739–49.
Jencks WP. Binding energy, specificity, and enzymic catalysis: the circe effect. Adv Enzymol Relat Areas Mol Biol. 1975;43:219–410.
Koshland DE. Application of a theory of enzyme specificity to protein synthesis. Proc Natl Acad Sci USA. 1958;44(2):98–104.
Koshland Jr DE. Crazy, but correct. Nature. 2004;432(7016):447.
Laidler KH. The influence of pressure on the rates of biological reactions. Arch Biochem. 1951;30(2):226–36.
Monod J, Wyman J, Changeux JP. On the nature of allosteric transitions: a plausible model. J Mol Biol. 1965;12:88–118.
Sullivan SM, Holyoak T. Enzymes with lid-gated active sites must operate by an induced fit mechanism instead of conformational selection. Proc Natl Acad Sci USA. 2008;105(37):13829–34.
Tsai CJ, Kumar S, Ma B, Nussinov R. Folding funnels, binding funnels, and protein function. Protein Sci. 1999a;8(6):1181–90.
Tsai CJ, Ma B, Nussinov R. Folding and binding cascades: shifts in energy landscapes. Proc Natl Acad Sci USA. 1999b;96(18):9970–2.
Voet D, Voet JG. Biochemistry. Hoboken: Wiley; 2004.
Download references
Author information
Authors and affiliations.
Department of Biochemistry and Molecular Biology, The University of Kansas Medical Center, 3901 Rainbow Boulevard, Kansas City, KS, 66160-7421, USA
Todd Holyoak
You can also search for this author in PubMed Google Scholar
Corresponding author
Correspondence to Todd Holyoak .

Editor information
Editors and affiliations.
Department of Biochemistry, University of Leicester, Leicester, UK
Gordon C. K. Roberts
Rights and permissions
Reprints and permissions
Copyright information
© 2013 European Biophysical Societies' Association (EBSA)
About this entry
Cite this entry.
Holyoak, T. (2013). Molecular Recognition: Lock-and-Key, Induced Fit, and Conformational Selection. In: Roberts, G.C.K. (eds) Encyclopedia of Biophysics. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-16712-6_468
Download citation
DOI : https://doi.org/10.1007/978-3-642-16712-6_468
Publisher Name : Springer, Berlin, Heidelberg
Print ISBN : 978-3-642-16711-9
Online ISBN : 978-3-642-16712-6
eBook Packages : Biomedical and Life Sciences Reference Module Biomedical and Life Sciences
Share this entry
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
- Publish with us
Policies and ethics
- Find a journal
- Track your research
The Lock and Key Theory: Understanding Enzyme Specificity and Catalysis
Mappa concettuale.
Modifica disponibile
The Lock and Key Theory, introduced by Emil Fischer, is a fundamental concept in biochemistry that explains enzyme specificity. It compares the enzyme's active site to a lock and the substrate to a key, illustrating how only the correct substrate can initiate a reaction. This theory is pivotal in understanding biochemical pathways, organic chemistry, and pharmaceuticals, influencing drug design by targeting enzyme active sites to treat diseases.
The Fundamentals of Enzyme Specificity: Lock and Key Theory
Exploring the Components and Dynamics of the Lock and Key Model
The role of the lock and key theory in organic chemistry and drug discovery, lock and key versus induced fit: diverse models of enzyme specificity, educational resources for grasping the lock and key theory, the enduring influence of the lock and key theory on biochemical science.
Mostra di più
Introduction to the Lock and Key Theory
Definition of the lock and key theory.
The Lock and Key Theory explains the specificity of enzyme action by comparing the enzyme's active site to a lock and the substrate to a key
Elements of the Lock and Key Theory
Enzyme and its active site
Enzymes are specialized proteins with a three-dimensional pocket, called the active site, designed to bind specific substrates
Substrates are molecules that bind to the enzyme's active site, triggering a specific reaction
Enzyme-substrate complex and product
The binding of the substrate to the active site forms an enzyme-substrate complex, which facilitates the conversion of the substrate into the product
Reaction sequence in the Lock and Key Theory
The Lock and Key Theory can be summarized by the reaction sequence \(E + S \rightarrow ES \rightarrow E + P\), where \(E\) represents the enzyme, \(S\) the substrate, \(ES\) the enzyme-substrate complex, and \(P\) the product
Applications of the Lock and Key Theory
Importance in biochemistry and pharmaceutical industry.
The Lock and Key Theory is a fundamental concept in biochemistry and is crucial in drug design for developing specific and effective pharmaceutical agents
Comparison with the Induced Fit Theory
The Induced Fit Theory complements the Lock and Key model by suggesting that the active site is dynamic and can adapt to accommodate slight variations in substrate structure
Educational resources for learning the Lock and Key Theory
Detailed diagrams and glossaries are valuable tools for understanding the step-by-step process of enzyme action and the key terms associated with the Lock and Key Theory
Significance of the Lock and Key Theory
Influence on our understanding of enzymatic function.
The Lock and Key Theory has greatly contributed to our understanding of enzymatic function and the complex chemical processes that sustain life
Practical applications in drug design
The Lock and Key Theory is essential in the development of enzyme inhibitors for treating diseases by disrupting pathological processes
Role in biochemistry education and research
The Lock and Key Theory remains a fundamental concept in biochemistry education and research, providing a starting point for exploring the complexities of enzyme behavior
Vuoi creare mappe dal tuo materiale?
Inserisci un testo, carica una foto o un audio su Algor. In pochi secondi Algorino lo trasformerà per te in mappa concettuale, riassunto e tanto altro!
Impara con le flashcards di Algor Education
Clicca sulla singola scheda per saperne di più sull'argomento.
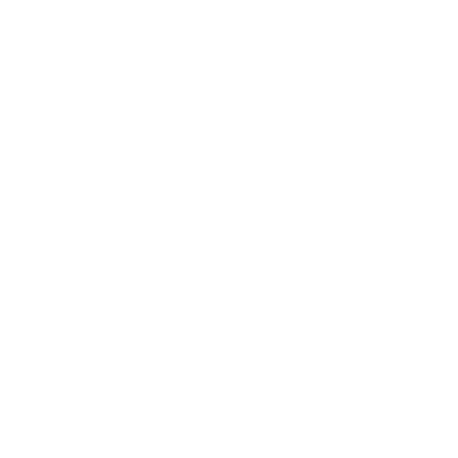

Enzymes, which are specialized ______, work as catalysts and their action is explained by the ______ and Key Theory.
proteins Lock

Describe the active site of an enzyme.
Three-dimensional pocket on enzyme surface; binds substrate with high specificity; involves non-covalent interactions like hydrogen bonds, ionic interactions, van der Waals forces.
Explain the enzyme-substrate complex.
Transient complex formed when substrate binds to enzyme's active site; facilitates substrate's conversion into product.
Summarize the reaction sequence in the Lock and Key Theory.
E + S -> ES -> E + P; E is enzyme, S is substrate, ES is enzyme-substrate complex, P is product; enzyme remains unchanged post-reaction.

The ______ and Key Theory is crucial for understanding enzyme catalysis in organic chemistry and drug design in the pharmaceutical industry.
Originator of Induced Fit Theory
Daniel Koshland proposed the Induced Fit Theory in 1958.
Characteristic of enzyme's active site in Induced Fit Theory
Active site is dynamic, molds around substrate upon binding.
Role of substrate structure variability in enzyme specificity
Enzyme adaptability allows accommodation of substrates with slight structural variations.
Educational materials like ______ and glossaries are crucial for grasping the ______ Theory.
detailed diagrams Lock and Key
Lock and Key Theory - Basic Concept
Theory where enzymes and substrates fit together precisely like a lock and key, explaining enzyme specificity.
Enzyme Specificity - Importance
Critical for enzymes to catalyze only the correct reactions, ensuring proper metabolic function.
Drug Design - Lock and Key Relevance
Lock and Key Theory guides creation of enzyme inhibitors that mimic substrates, blocking unwanted reactions in disease.
Ecco un elenco delle domande più frequenti su questo argomento
Who introduced the lock and key theory and what does it explain about enzymes, what are the key components involved in the lock and key theory, how does the lock and key theory aid in the development of new pharmaceuticals, what is the difference between the lock and key and the induced fit theories of enzyme specificity, what educational tools are recommended for understanding the lock and key theory, despite advancements in biochemistry, why is the lock and key theory still significant, contenuti simili, esplora altre mappe su argomenti simili.
Brain Development and Its Impact on Functioning
Amino Acids and Proteins
Messenger RNA and Protein Synthesis
Protein Structure and Function
Ribosomal RNA and its Role in Protein Synthesis
Enzymes: Biological Catalysts for Life
Glycolipids: Structure, Function, and Importance

Non trovi quello che cercavi?
Cerca un argomento inserendo una frase o una parola chiave
Cosa ne pensi di noi?
Il tuo nome
La tua email
Structural Biochemistry/Protein function/Lock and Key
In the Lock and Key Model, first presented by Emil Fisher, the lock represents an enzyme and the key represents a substrate. It is assumed that both the enzyme and substrate have fixed conformations that lead to an easy fit. Because the enzyme and the substrate are at a close distance with weak attraction, the substrate must need a matching shape and fit to join together. At the active sites, the enzyme has a specific geometric shape and orientation that a complementary substrate fits into perfectly. The theory behind the Lock and Key model involves the complementarity between the shapes of the enzyme and the substrate. Their complementary shapes make them fit perfectly into each other like a lock and a key. According to this theory, the enzyme and substrate shape do not influence each other because they are already in a predetermined perfectly complementary shape. As a result, the substrate will be stabilized. This theory was replaced by the induced fit model which takes into account the flexibility of enzymes and the influence the substrate has on the shape of the enzyme in order to form a good fit.

The active site is the binding site for catalytic and inhibition reaction of the enzyme and the substrate; structure of active site and its chemical characteristic are of specificity for binding of substrate and enzyme. Three models of enzyme-substrate binding are the lock-and-key model, the induced fit model, and the transition-state model. The lock-and-key model assumes that active site of enzyme is good fit for substrate that does not require change of structure of enzyme after enzyme binds substrate.
- Book:Structural Biochemistry
Navigation menu

IMAGES
VIDEO
COMMENTS
Lock-and-key model Definition. Lock-and-key model is a model for enzyme-substrate interaction suggesting that the enzyme and the substrate possess specific complementary geometric shapes that fit exactly into one another. In this model, enzymes are depicted as highly specific.
Learn how enzymes catalyze chemical reactions by lowering the activation energy using the lock and key model. Find out the limitations of this model and the induced fit model that explains the enzyme-substrate complex formation.
This article reviews the history and evolution of three models of molecular recognition: lock-and-key, induced fit, and conformational selection. It explains the key features, assumptions, and applications of each model, and how they are related to enzyme catalysis and protein interactions.
Learn about the lock-key model and its variations for enzyme catalysis, with examples and references. The keyhole-lock-key model considers the role of tunnels and channels in substrate binding and product release.
In fact, an early model describing the formation of the enzyme-substrate complex was called the lock-and-key model (Figure \(\PageIndex{2}\)). This model portrayed the enzyme as conformationally rigid and able to bond only to substrates that exactly fit the active site.
Learn how enzymes and substrates interact in the lock-and-key model of enzyme action. See how the induced-fit model refines the original model and how enzymes have substrate specificity.
Learn how the Lock and Key Theory, proposed by Emil Fischer, explains the specificity of enzyme action. The theory compares the enzyme's active site to a lock and the substrate to a key, illustrating how only the correct substrate can initiate a reaction.
The lock and key model, proposed by Emil Fisher, assumes that enzymes and substrates have fixed conformations that fit perfectly like a lock and a key. This model was later replaced by the induced fit model, which considers the flexibility and interaction of enzymes and substrates.
This book celebrates the centenary of Emil Fischer's lock-and-key hypothesis, which postulated that molecules can fit together like keys and locks. It explores the applications and implications of this concept in biology, catalysis, molecular devices, and the origin of life.
This chapter contains sections titled: Introduction. Classical Locks. Catenanes. Safety Locks. Magnetic Locks. Carbohydrate-Lectin Interactions: Supracellular Chemistry. Conclusion.